HOWTO Execute a Launch using NERSC
Contents
Introduction
This page gives some instructions on executing a launch at NERSC. Note that some steps must be completed to make sure things are set up at Cori and Globus prior to submitting any jobs.
The following is based on steps used to do RunPeriod-2018-01 monitoring launch ver 18 using swif2.
Quick Start
- ssh from gxproj4 account to cori.nersc.gov to make sure passwordless login works
- login to Globus and make sure both endpoints are active ("NERSC DTN", "jlab#scidtn1")
- Make directory in gxproj4 for launch, checkout launch scripts, and modify launch/launch_nersc.py
- mkdir ~gxproj4/NERSC/2018.10.05.offmon_ver18
- cd ~gxproj4/NERSC/2018.10.05.offmon_ver18
- svn co https://halldsvn.jlab.org/repos/trunk/scripts/monitoring/launch
- svn co https://halldsvn.jlab.org/repos/trunk/scripts/monitoring/hdswif2
- Run launch_nersc.py in test mode with VERBOSE=3 and for 1 file of 1 run and check swif2 command carefully
- Commit any changes to launch directory scripts to repository
- ssh to cori.nersc.gov and update launch directory
- cd projectdir_JLab/launch
- svn update
- Make sure enough scratch disk space is available on cori (use myquota)
- Back at ifarm, turn test mode off, and point output to halld-scratch in launch_nersc.py and run test job
- IF job run successfully:
- turn test mode back on
- point output back to final destination (not halld-scratch)
- set verbose to 1
- modify numbers to process all files desired for launch
- run launch_nersc.py and confirm everything looks right
- turn off test mode and initiate launch
- Set up updating job monitoring plots
- ssh into gxproj4 on ifarm in terminal that can be left up for long periods (e.g. desktop)
- cd to project directory ( cd ~gxproj4/NERSC/2018.10.05.offmon_ver18
- checkout job monitoring scripts: svn co https://halldsvn.jlab.org/repos/trunk/scripts/monitoring/hdswif2
- cd hdswif2
- Modify auto_run.sh to reflect correct swif2 workflow name
- Modify uploaddir in regenerate_plots.csh to point to correct directory to upload files to
- Add section to regenerate_plots.csh for new workflow
- ssh to gxproj5@ifarm and:
- mkdir the uploaddir
- modify /group/halld/www/halldweb/html/data_monitoring/launch_analysis/index.html to include new launch campaign
- log back out to the previous gxproj4 shell
- setup environment for ROOT: source /apps/root/PRO/setroot_CUE.csh
- run: ./auto_run.sh
NERSC Account
To run jobs at NERSC you need to get a user account there. This account will need to be associated with a repository which is what they call a project that has some resources allocated to it. At this time, the GlueX project is m3120. You can find instructions for applying for an account here: http://www.nersc.gov/users/accounts/user-accounts/get-a-nersc-account.
Setting up SSH
Swif2 will access Cori at NERSC via passwordless login. To set this up, you’ll need a RSA key with empty passphrase and the public key installed on the NERSC account to be used. Chris’ instruction for this are:
(2-a) See http://www.nersc.gov/users/connecting-to-nersc/connecting-with-ssh (2-b) As the user who owns the workflow, generate an ssh key as specified, supply no passphrase (2-c) log in to nim.nersc.gov, and under "My ssh keys" add the public key you generated in (2-b) (2-d) verify that you can login to cori.nersc.gov without a password after logging in to ifarm as the workflow user.
For this to actually work, you must make sure that the key created above is what is used when authenticating. I created a dedicated key pair with the names ~/.ssh/id_rsa_nersc and ~/.ssh/id_rsa_nersc.pub . There are multiple ways to use this key (ssh-agent, using the ‘-i’ option with the ssh command,...). The best way to use it with swif2 though is to specify the key in the ~/.ssh/config file. There you can specify that the special key is used when logging into cori.nersc.gov and furthermore which username to use when logging in. This last part is important since I needed it to log into my davidl account from the gxproj4 account at jlab.
Here are the lines that need to be in the ~/.ssh/config file:
# The following is to allow passwordless login to
# cori.nersc.gov without having to run an agent or
# explicitly give the key on the ssh command.
# n.b. this will login as davidl and not in some
# group account at nersc (since none currently
# exists)
# 7/18/2018 DL
Host cori cori.nersc.gov
IdentityFile ~/.ssh/id_rsa_nersc
User davidl
Files and directories on Cori at NERSC
When jobs are run at NERSC that will need access to a couple of files from the launch directory where we keep GlueX farm submission scripts and files. This is kept in our subversion repository at JLab. When a job is started at NERSC it will look for this directory in the project directory that swif2 is using for the workflow (see next section). The launch directory should already be checked out there but for completeness, here is how you would do it:
cd /global/project/projectdirs/m3120 svn co https://halldsvn.jlab.org/repos/trunk/scripts/monitoring/launch
If the directory is there, make sure it is up to date:
cd /global/project/projectdirs/m3120 svn update
The most important files are the script_nersc.py script and the jana_offmon_nersc.config (jana_recon_nersc.config) files. This first is what is actually run inside of the container when the job wakes up. The second specifies the plugins and other settings. For the most part, the "nersc" versions of the jana config files should be kept in alignment with the JLab versions. One notable difference is that the NERSC jobs are always run on whole nodes so NTHREADS is always set to "Ncores" whereas at JLab they are usually set to 24.
*** IMPORTANT *** At this point you should check the available scratch disk space for the account you will use to run the launch using the myquota command on Cori. If sufficient space is not available (i.e. 27GB x MAX_CONCURRENT_JOBS) then clear it out now.
Globus Endpoint Authentication
Submitting jobs to swif2
The offsite jobs at NERSC are managed from the gxproj4 account. This is a group account with access limited to certain users. Your ssh key must be added to the account by an existing member. Contact the software group to request access.
Generally, one would log into an appropriate computer with:
ssh gxproj4@ifarm
The following are some steps needed to create a workflow and submit jobs.
Create a new workflow
This step can actually be skipped since the script in the next step will automatically create the workflow with the correct name and parameters if it does not already exist. These are instructions in case you want/need to create the workflow yourself.
The workflow name follows a convention based on the type of launch, run period, version, and optional extra qualifiers. Here is the command used to create the workflow for offline monitoring launch ver18 for RunPeriod-2018-01:
swif2 create -workflow offmon_2018-01_ver18 -max-concurrent 2000 -site nersc/cori -site-storage nersc:m3120
The -max-concurrent 2000 option tells swif2 to limit the number of dispatched jobs to no more than 2000. The primary concern here is in scratch disk space at NERSC. If each input file is 20GB and produces 7GB of output then the we need 27GB * 2000 = 54 TB of free scratch disk space. If multiple launches are running at the same time and using the same account's scratch disk then it is up to you to make sure the sum of requirements does not exceed the quota. At this point in time we have a quota of 60TB of scratch space, though they have claimed that they will revisit that at the beginning of the year.
The -site nersc/cori is required at the moment and is the only allowed option for "site".
The -site-storage nersc:m3120 is used to specify which NERSC project assigned disk space to use. At this point, swif2 has been changed to use scratch disk space assigned to the personal account being used to run the jobs so I believe this is being ignored.
Prepare working directory at JLab
Create a working directory in the gxproj4 account and checkout the launch scripts. This is done so that the scripts can be modified for the specific launch in case some tweaks are needed. Changes should eventually be pushed back into the repository, but having a dedicated directory for the launch can help with managing things, especially when there are multiple launches.
mkdir ~gxproj4/NERSC/2018.10.05.offmon_ver18 cd ~gxproj4/NERSC/2018.10.05.offmon_ver18 svn co https://halldsvn.jlab.org/repos/trunk/scripts/monitoring/launch
Configure the parameters for the launch
At this time the parameters used for a NERSC launch are specified in the launch_nersc.py script. This is slightly different for jobs run at JLab which use the launch.py script that reads the configuration from a separate file. At some point the NERSC system should be brought more into alignment with that, but for now, this is how it is.
Edit the file launch/launch_nersc.py to adjust all of the settings at the top to be consistent with the current launch. All of the parameters are at the top of the file in a well marked section. Here is an explanation of the parameters:
| Parameter | Description |
|---|---|
| TESTMODE | Set this to "True" so the script can be tested without actually submitting any jobs. When finally ready to actually submit jobs to swif2, set it to "False" |
| VERBOSE | Default is 1. Set to zero for minimal messages or 3 for all messages |
| RUNPERIOD | e.g. "2018-01" |
| LAUNCHTYPE | either "offmon" or "recon" |
| VER | Version of this particular type of launch |
| WORKFLOW | This will be set automatically based on other values. Only change this if there default name is not appropriate |
| NAME | Similar to above, this is automatically set. It is used to set the job names. |
| RCDB_QUERY | If specific runs are not set in RUNS (see below) then this is used to query the RCDB for runs in the specified range that should be processed. |
| RUNS | Normally this is set to an empty array and the list of runs obtained from the RCDB. This can be set to a specific run list and only those will be processed. |
| MINRUN | If RUNS is an empty set, this is used along with MAXRUN and RCDB_QUERY to extract the list of runs to process from the RCDB. Note that this should be in a range consistent with RUNPERIOD. No check is made in this script that ensures this otherwise. |
| MAXRUN | See MINRUN above. |
| MINFILENO | Minimum file number to process for each run. Normally this is set to 0. See MAXFILENO below for more details. |
| MAXFILENO | Maximum file number to process for each run. If doing a monitoring launch then this would normally be set to 4 so that files 000-004 are processed. Set this to a large number like 10000 to process all files in each run. The RCDB will be queried for each run to so that only files that actually exist in the specified range are submitted as jobs. |
| MAX_CONCURRENT_JOBS | This is a limit set on the swif2 workflow for how many jobs can be in-flight at once. This can only be specified when the workflow is created. If the workflow was created outside of this script then this will do nothing. |
| PROJECT | The NERSC project. Use 'm3120' for the GlueX allocation |
| TIMELIMIT | Maximum time job may run. If a job runs longer than this it will be killed. Keep in mind that jobs run on KNL take about 2.4 times as long to run as on Haswell. (See NODETYPE below.) |
| QOS | Should be 'debug', 'regular', or 'premium'. Usually you will just want 'regular' |
| NODETYPE | 'haswell' or 'knl'. Jobs will take about 2.4 times longer to run on knl as haswell and the charge rate is 20% more per node hour. However, there are only 2k haswell nodes and 9k knl nodes and much more demand for haswell. Note that the setting of TIMELIMIT should be adjusted based on this. |
| IMAGE | Shifter image. Actually, this is the same name as the Docker image used to create the Shifter image. Note that this image will need to already exist in Shifter since it will not be pulled in automatically. The image used up to now is 'docker:markito3/gluex_docker_devel' |
| RECONVERSION | This the version of the reconstruction code that should be used. The executables will be read from CVMFS from a directory that mirrors the directory /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr. This value should be a directory relative to that such as: halld_recon/halld_recon-recon-ver03.2 |
| SCRIPTFILE | The script to run inside the container for the job. The container will mount the launch directory as /launch in the container so this should normally be set to '/launch/script_nersc.sh |
| CONFIG | The jana config file to use. This is set automatically but can be overridden if really needed. As with the SCRIPTFILE, this should access the file via the launch directory mounted in the container as /launch |
| OUTPUTTOP | Location on the JLab queue to copy files back to. Usually, this will start with mss:/mss/halld/ indicating a place on the tape system. This specifies the top-level directory and the launch_nersc.py script will specify the subdirectories where individual output files should be copied. |
| RCDB_HOST | Were the launch_nersc.py script should access the RCDB to extract the run numbers to be used for this launch. |
| RCDB_USER | User to connect to the RCDB as. (See RCDB_HOST above.) |
| RCDB | This is used internally in the script and should always be set to None |
Archiving Launch Parameters
- Edit the appropriate html file in the run period's /group/halld/www/halldweb/html/data_monitoring/launch_analysis/ subdirectory (e.g. /group/halld/www/halldweb/html/data_monitoring/launch_analysis/2018_01/2018_01.html)
There may be two places to add the current launch
- Copy JANA config file to archive directory:
cd ~gxproj4/NERSC/2018.10.05.offmon_ver18/launch cp jana_offmon_nersc.config /group/halld/data_monitoring/run_conditions/RunPeriod-2018-01/jana_offmon_2018_01_ver18.config
- Copy halld_recon version file to archive directory:
cp /group/halld/www/halldweb/html/dist/version_3.2.xml /group/halld/data_monitoring/run_conditions/RunPeriod-2018-01/version_offmon_2018_01_ver18.xml
Troubleshooting
There are many places and ways that jobs can fail and it can be difficult to find information since it is dispersed over several systems. Here are some tips for tracking down issues.
SWIF2
SWIF2 is the starting and ending point for each job so it is an important first step. Unfortunately, with thousands of jobs, it is not usually practical to dump information to the screen scan it for the one you're interested in.
Finding the swif2 jobid: swif2 show-job -workflow offmon_2018-01_ver18 -name GLUEX_offmon_041261_001
Listing problem jobs: swif2 status -problems -workflow offmon_2018-01_ver18
Setting Time Limit: swif2 modify-jobs -workflow offmon_2018-01_ver18 -time set 10h -names GLUEX_offmon_040902_004
n.b. The normal swif2 job submission sets a time limit via a sbatch option. Swif2 passes this along, but does not record it as a swif2 option. This means using the swif2 "-time add" or "-time mult" options will not work unless you have run "-time set" on the job already. Once the time limit is in swif2, it will automatically add it's time limit to the sbatch command when the job is retried.
n.b. Modifying the job seems to automatically retry it so you should not need to run "swif2 retry-jobs".
Here is a script for finding the SLURM_TIMEOUT jobs in the owrkflow and setting them to a new timelimit
#!/usr/bin/env python
import json
import subprocess
# Create list of problem jobs with:
#
# swif2 status -problems -workflow offmon_2018-01_ver18 -display json > problem_jobs.json
problem_type = 'SLURM_TIMEOUT'
with open('problem_jobs.json') as f:
data = json.load(f)
cmd = ['swif2', 'modify-jobs', '-workflow', 'offmon_2018-01_ver18', '-time', 'set', '10h', '-names']
for job in data:
if job['job_attempt_problem'] == problem_type:
cmd.append(job['job_name'])
print ' '.join(cmd)
subprocess.call(cmd)
Output File on Cache seems small or is corrupted
Looking at the file sizes on the cache disk, the 001 REST file seems small:
ifarm1401:gxproj4:~> ls -l /cache/halld/offline_monitoring/RunPeriod-2018-01/ver18/REST/041261/ total 38600901 -rw-rw-r-- 1 davidl halld-2 4799039221 Oct 10 06:52 dana_rest_041261_000.hddm -rw-rw-r-- 1 davidl halld-2 413039444 Oct 9 04:39 dana_rest_041261_001.hddm -rw-rw-r-- 1 davidl halld-2 4907285495 Oct 10 06:16 dana_rest_041261_003.hddm -rw-rw-r-- 1 davidl halld-2 4901814991 Oct 10 03:28 dana_rest_041261_004.hddm -rw-rw-r-- 1 davidl halld-2 4897558125 Oct 10 02:05 dana_rest_041261_005.hddm -rw-rw-r-- 1 davidl halld-2 4889533411 Oct 10 03:34 dana_rest_041261_006.hddm -rw-rw-r-- 1 davidl halld-2 4883590490 Oct 10 06:09 dana_rest_041261_007.hddm -rw-rw-r-- 1 davidl halld-2 4879139499 Oct 10 03:35 dana_rest_041261_008.hddm -rw-rw-r-- 1 davidl halld-2 4876425372 Oct 10 09:02 dana_rest_041261_009.hddm
1. Make sure that the file is still not being transferred by checking the modification time.
- If it is less than 1 hr, then give it a little more time. Files do not necessarily get processed in order so this may just be the last one
2. Check the status of the job in swif2
- Use the run number, file number, and type (offmon, recon, ...) to get the job status by name (click expand on right of next line to see output of command)
ifarm1401:gxproj4:~> swif2 show-job -workflow offmon_2018-01_ver18 -name GLUEX_OFFMON_041261_001
job_id = 7878 job_name = GLUEX_offmon_041261_001 workflow_name = offmon_2018-01_ver18 workflow_user = gxproj4 job_status = done job_attempt_status = done num_attempts = 1 site_job_command = /global/project/projectdirs/m3120/launch/run_shifter.sh--module=cvmfs--/launch/script_nersc.sh/launch/jana_offmon_nersc.confighalld_recon/halld_recon-recon-ver03.2412611 site_job_batch_flags = -Am3120--volume="/global/project/projectdirs/m3120/launch:/launch"--image=docker:markito3/gluex_docker_devel--time=9:00:00--nodes=1--tasks-per-node=1--cpus-per-task=64--qos=regular-Cknl-Lproject ... job_attempt_id = 10658 site_job_id = 7893 job_attempt_status = done slurm_id = 15513434 job_attempt_cleanup = done site_job_id = 7893 job_id = 7878 site_id = 1 site_job_command = /global/project/projectdirs/m3120/launch/run_shifter.sh--module=cvmfs--/launch/script_nersc.sh/launch/jana_offmon_nersc.confighalld_recon/halld_recon-recon-ver03.2412611 site_job_batch_flags = -Am3120--volume="/global/project/projectdirs/m3120/launch:/launch"--image=docker:markito3/gluex_docker_devel--time=9:00:00--nodes=1--tasks-per-node=1--cpus-per-task=64--qos=regular-Cknl-Lproject site_id = 1 jobid = 15513434 jobstep = batch avecpu = 00:00:00 avediskread = 1575.82M averss = 3955K avevmsize = 21408K cputime = 11-11:55:44 elapsed = 01:00:52 end = 2018-10-07 14:19:03.0 exitcode = 0 start = 2018-10-07 13:18:11.0 state = COMPLETED maxdiskread = 9850.38M maxdiskwrite = 2003.68M maxpages = 198K maxrss = 34648872K maxvmsize = 45004068K exitsignal = 0 avediskwrite = 1571.86M
You'll notice in the above command that the state is "COMPLETED". The total time though was just over an hour and this job should have taken more than 8 hours to run. Thus, something happened, but swif2 did not recognize it as an error.
3. Check the status of the job in SLURM
- Using the "slurm_id" from the swif2 output above, log into cori.nersc.gov and check the status. Note that this command assumes the job is finished
davidl@cori07:~> sacct -j 15513434
JobID JobName Partition Account AllocCPUS State ExitCode ------------ ---------- ---------- ---------- ---------- ---------- -------- 15513434 GLUEX_off+ regular m3120 272 COMPLETED 0:0 15513434.ba+ batch m3120 272 COMPLETED 0:0 15513434.ex+ extern m3120 272 COMPLETED 0:0
Here, slurm also thinks everything went fine and claims an exit code of 0.
4. Look at the job output
- Since this job completed successfully according to swif2, the output files have likely already been deleted from NERSC so we'll need to access the copy transferred to JLab. First look for it on the cache disk:
ls /cache/halld/offline_monitoring/RunPeriod-2018-01/ver18/job_info/041261/job_info_041261_001.tgz
If the file is not there then you'll have to request it from the tape library using jcache
- The stdout and stderr files are inside the .tgz file. Unpack it and have a look:
ifarm1401:gxproj4:~> tar xzf /cache/halld/offline_monitoring/RunPeriod-2018-01/ver18/job_info/041261/job_info_041261_001.tgz ifarm1401:gxproj4:~> ls job_info_041261_001 cpuinfo.out env.out hostname.out std.err std.out top.out
- It is worth knowing what the normal output of a job looks like since there are often warnings or other messages that look like problems, but actually aren't. Don't be misled by those. Taking a look at the std.err files shows:
ifarm1401:gxproj4:~>tail -n 200 job_info_041261_001/std.err
... Generating stack trace... JANA E 0x00002aaabdd30ba4 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabc7d2ec8 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcacc386 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcac3520 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcac4362 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcac1220 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcb2959f in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcb2eedb in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcb2f0f2 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcb2f22f in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcaef167 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabcb35db3 in <unknown> from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabc6cfe26 in cling::LookupHelper::findScope(llvm::StringRef, cling::LookupHelper::DiagSetting, clang::Type const**, bool) const + 0x486 from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaabc652674 in TCling::CheckClassInfo(char const*, bool, bool) at /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/core/meta/src/TCling.cxx:3456 from /group/halld/Software/builds/Linux_CentOS7-x86_64- gcc4.8.5-cntr/root/root-6.08.06/lib/libCling.so 0x00002aaaab52f68e in TClass::GetClass(char const*, bool, bool) at /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/core/meta/src/TClass.cxx:3039 (discriminator 1) from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libCore.so 0x00002aaaab9a3f9b in TGenCollectionProxy::Value::Value(std::basic_string<char, std::char_traits<char>, std::allocator<char> > const&, bool) at /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/build_dir/include/TClassRef.h:62 from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libRIO.so 0x00002aaaab983451 in TEmulatedCollectionProxy::InitializeEx(bool) at /usr/include/c++/4.8.2/bits/atomic_base.h:783 (discriminator 1) from /group/halld/Software/builds/Linux_CentOS7-x86_64-gcc4.8.5-cntr/root/root-6.08.06/lib/libRIO.so ...
- So this looks like an error occurred in ROOT, but it somehow managed to exit with a clean exit code. The std.out file doesn't help since it looks like it just suddenly stopped in the middle of processing.
5. Resurrect the job
- This particular issue looks like it is probably some bug exposed through a rare race condition. (Rare since we don't see large fractions of jobs doing this.) The best thing to do in this case would be to re-try the job. Since swif2 thinks the job finished OK, we have to tell it to "resurrect" the job. This is a feature I don't see in the swif2 help, but Chris told me about it.
- n.b. The resurrected job will copy files with the same name back to the write-through cache at JLab which will eventually replace the version on tape.
ifarm1401:gxproj4:~> swif2 retry-jobs -resurrect -workflow offmon_2018-01_ver18 7878 Found 1 matching jobs Resurrecting 1 successful job
6. Find similar bad jobs
- This problem was a little more insidious because no error code was returned and all output files were at least partially produced. This meant swif2 could not tell us there was a problem. Once one problem like this is discovered, you have figure out a way to look for other similar ones. Here, the easiest thing to do is look for small REST files which is what tipped us off to start with. Well, small REST files relative to the input EVIO file size. Each run will end with a partial file and therefore have a small file. For this, I wrote a script to extract the file sizes from the mss stub files.
Script to get file sizes of all REST files and their corresponding EVIO raw data files
#!/usr/bin/env python
import glob
import linecache
RESTDIR = '/mss/halld/offline_monitoring/RunPeriod-2018-01/ver18/REST'
RAWDIR = '/mss/halld/RunPeriod-2018-01/rawdata'
for f in glob.glob( RESTDIR+'/*/dana_rest_*.hddm'):
# Get size of REST file
theline = linecache.getline(f, 3) # size is 3rd line in file
fsize = theline.split('=')[1].strip()
# Get size of EVIO raw data file
run_split = f[-15:-5] # extract run/file number from REST file name
run = run_split[0:6 ] # Extract run number
split = run_split[8:11] # Extract split number
fevio = RAWDIR + '/Run' + run + '/hd_rawdata_' + run_split + '.evio'
theline = linecache.getline(fevio, 3) # size is 3rd line in file
feviosize = theline.split('=')[1].strip()
ratio = float(fsize)/float(feviosize)
print '%s %s %s %s %f' % (run, split, fsize, feviosize, ratio)
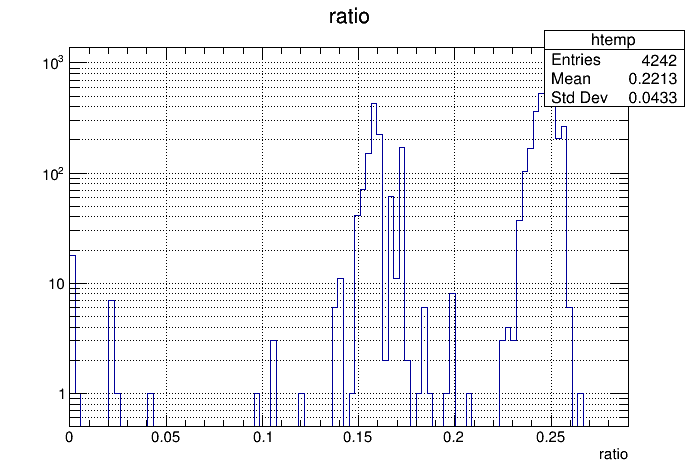
- Run this script and capture its output to a text file. This makes it easy to read into ROOT. The reason for reading it into ROOT is actually apparent when looking at the ratio of REST to EVIO file sizes. The distribution is bimodal so we need to be careful how to cut on what is "bad" (see plot below). Some of these are clearly bad, but what is less clear are the ones with a ratio between 0.18 and 0.21. These must be looked at to see if they need to be re-run or not. This requires going back to the job_info files as described in step 4 above.