Difference between revisions of "CDC algo tuning"
From GlueXWiki
| Line 30: | Line 30: | ||
{| border="0" cellpadding="2" | {| border="0" cellpadding="2" | ||
|+ High and low thresholds found using upsampled data, no interpolation | |+ High and low thresholds found using upsampled data, no interpolation | ||
| − | |[[Image: | + | |[[Image:cdc_run_31942_track127b_np16_thr_5_4_1.png|thumb|x150px|Thresholds 5-4-1 pedlead 16 tz 7 tfix 1000]] |
|} | |} | ||
Revision as of 14:13, 4 December 2013
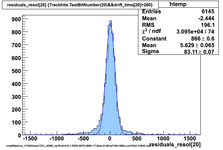
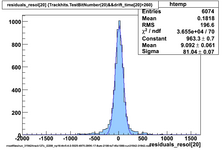
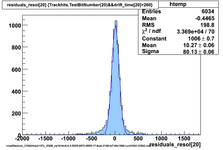
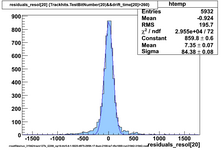
Using prototype data from CDC_50_50 (run 31924) with offline analysis
50/50 Ar/CO2 and cosmics, 2100V, prototype horizontal
Original code, many samples upsampled
- Find hit threshold crossing
- Step back <pedlead> points to find new pedestal
- Search forward to find high threshold crossing
- Search backward to find low threshold crossing
- Project through both thresholds to find pedestal crossing time
Fewer samples upsampled
- Find hit threshold crossing
- Upsample region around threshold crossing, with threshold crossing sample in set position
- Step back <pedlead> points to find new pedestal
- Search forward to find high threshold crossing
- Search backward to find low threshold crossing
- Project through both thresholds to find pedestal crossing time
Ideas
- Check whether interpolating threshold xings makes any difference
- ? Pass in hit pedestal to upsampling module, to find initial hit threshold xing better
- Or interpolate around initial hit threshold xing and pass in number of 1/5 sample time?
- Try reverting to one threshold crossing instead of the extrapolation